Experts Matter. Find Yours.
Connect for media, speaking, professional opportunities & more.

• Has potential to help geneticists investigate vital issues such as antibacterial resistance • Will untangle the genetic components shared due to common ancestry from the ones shared due to evolution • The work is result of a four-year international collaboration. Aston University has worked with international partners to develop a software package to help scientists answer key questions about genetic factors associated with shared characteristics among different species. Called CALANGO (comparative analysis with annotation-based genomic components), it has the potential to help geneticists investigate vital issues such as antibacterial resistance and improvement of agricultural crops. This work CALANGO: a phylogeny-aware comparative genomics tool for discovering quantitative genotype-phenotype associations across species has been published in the journal Patterns. It is the result of a four year collaboration between Aston University, the Federal University of Minas Gerais in Brazil and other partners in Brazil, Norway and the US. Similarities between species may arise either from shared ancestry (homology) or from shared evolutionary pressures (convergent evolution). For example, ravens, pigeons and bats can all fly, but the first two are birds whereas bats are mammals. This means that the biology of flight in ravens and pigeons is likely to share genetic aspects due to their common ancestry. Both species are able to fly nowadays because their last common ancestor – an ancestor bird - was also a flying organism. In contrast, bats have the ability to fly via potentially different genes than the ones in birds, since the last common ancestor of birds and mammals was not a flying animal. Untangling the genetic components shared due to common ancestry from the ones shared due to common evolutionary pressures requires sophisticated statistical models that take common ancestry into account. So far, this has been an obstacle for scientists who want to understand the emergence of complex traits across different species, mainly due to the lack of proper frameworks to investigate these associations. The new software has been designed to effectively incorporate vast amounts of genomic, evolutionary and functional annotation data to explore the genetic mechanisms which underly similar characteristics between different species sharing common ancestors. Although the statistical models used in the tool are not new, it is the first time they have been combined to extract novel biological insights from genomic data. The technique has the potential to be applied to many different areas of research, allowing scientists to analyse massive amounts of open-source genetic data belonging to thousands of organisms in more depth. Dr Felipe Campelo from the Department of Computer Science in the College of Engineering and Physical Sciences at Aston University, said: “There are many exciting examples of how this tool can be applied to solve major problems facing us today. These include exploring the co-evolution of bacteria and bacteriophages and unveiling factors associated with plant size, with direct implications for both agriculture and ecology.” “Further potential applications include supporting the investigation of bacterial resistance to antibiotics, and of the yield of plant and animal species of economic importance.” The corresponding author of the study, Dr Francisco Pereira Lobo from the Department of Genetics, Ecology and Evolution at the Federal University of Minas Gerais in Brazil, said: “Most genetic and phenotypic variations occur between different species, rather than within them. Our newly developed tool allows the generation of testable hypotheses about genotype-phenotype associations across multiple species that enable the prioritisation of targets for later experimental characterization.” For more details about studying computer since at Aston University visit https://www.aston.ac.uk/eps/informatics-and-digital-engineering/computer-science

New research suggests model organisms may have evolved too far
A research team from Aston University and the Universities of Birmingham and Nottingham suggest model organisms evolved over 100 years may no longer be fit for purpose They found the bacterial strain Escherichia coli K-12 has been repeatedly cultured and mutated, resulting in many genetic changes The study has just been published in Microbial Genomics A research team from Aston University has found that the model organism used in laboratories for the past 100 years has evolved so extensively that it may no longer be fit for purpose. According to a new study, published in Microbial Genomics, the bacterial strain Escherichia coli K-12 has been repeatedly cultured and mutated, resulting in an organism that carries many genetic changes compared to the original isolated bacteria. The research team, from Aston University, and the Universities of Birmingham and Nottingham, made their discovery after re-examining the early preserved samples and looking at the base sequence of their DNA. They found a large number of differences at the DNA sequence level, and the differences are bigger when they examined currently used stocks that derived from the original samples. The work underscores the dangers of using one strain as a sole model. It also confirms that bacterial sequences evolve over short time scales and provides a fascinating insight into the first baby steps of molecular microbiology. Lead author Dr Doug Browning, of the School of Biosciences at Aston University, said: “The past 10 years have seen a massive amount of bacterial genome sequencing and the picture that is emerging is that bacterial genomes change very fast. This was unimaginable 100 years ago, and, of course, this is why folk back then were quick to adopt the K-12 strain as the model for everything.” The strain of bacteria in the study was originally isolated in 1922 from the faeces of a recuperating diptheria patient at Stanford University, in California. The strain was preserved and over time it, and many derivatives, were distributed to research laboratories around the world for use by researchers looking to understand the workings of living cells at the molecular level. While the number of genetic variations which have appeared in the intervening decades may sound alarm bells in some research areas, for others it may represent new research opportunities. Co-author Steve Busby, of the Institute of Microbiology and Infection at the University of Birmingham, said: “Actually the diversity that all this generates can add a new dimension to our understanding. It’s often true that things are seldom as they seem, and particularly so if you only study one strain.”
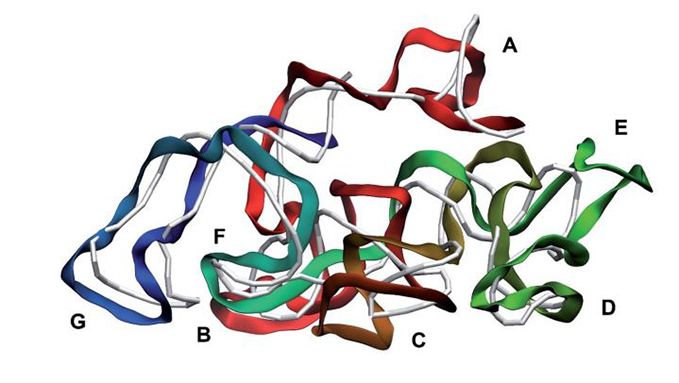
First ever computer reconstruction of a virus, including its complete native genome Will open way for investigating biological processes which can’t currently be fully examined because the genome is missing Could lead the way to research into an alternative to antibiotics. An Aston University researcher has created the first ever computer reconstruction of a virus, including its complete native genome. Although other researchers have created similar reconstructions, this is the first to replicate the exact chemical and 3D structure of a ‘live’ virus. The breakthrough could lead the way to research into an alternative to antibiotics, reducing the threat of anti-bacterial resistance. The research Reconstruction and validation of entire virus model with complete genome from mixed resolution cryo-EM density by Dr Dmitry Nerukh, from the Department of Mathematics in the College of Engineering and Physical Sciences at Aston University is published in the journal Faraday Discussions. The research was conducted using existing data of virus structures measured via cryo-Electron Microscopy (cryo-EM), and computational modelling which took almost three years despite using supercomputers in the UK and Japan. The breakthrough will open the way for biologists to investigate biological processes which can’t currently be fully examined because the genome is missing in the virus model. This includes finding out how a bacteriophage, which is a type of virus that infects bacteria, kills a specific disease-causing bacterium. At the moment it is not known how this happens, but this new method of creating more accurate models will open up further research into using bacteriophage to kill specific life-threatening bacteria. This could lead to more targeted treatment of illnesses which are currently treated by antibiotics, and therefore help to tackle the increasing threat to humans of antibiotic resistance. Dr Nerukh said: “Up till now no one else had been able to build a native genome model of an entire virus at such detailed (atomistic) level. “The ability to study the genome within a virus more clearly is incredibly important. Without the genome it has been impossible to know exactly how a bacteriophage infects a bacterium. “This development will now allow help virologists answer questions which previously they couldn’t answer. “This could lead to targeted treatments to kill bacteria which are dangerous to humans, and to reduce the global problem of antibiotic-resistant bacteria which are over time becoming more and more serious.” The team’s approach to the modelling has many other potential applications. One of these is creating computational reconstructions to assist cryo-Electron Microscopy – a technique used to examine life-forms cooled to an extreme temperature.

#Expert Research: New National Science Foundation and NASA-Funded Research Investigates Martian Soil
Studies have shown crops can grow in simulated Martian regolith. But that faux material, which is similar to soil, lacks the toxic perchlorates that makes plant growth in real Red Planet regolith virtually impossible. New research involving Florida Tech is examining how to make the soil on Mars useful for farming. Andrew Palmer, co-investigator and ocean engineering and marine sciences associate professor, along with Anca Delgado, principal investigator and faculty member at Arizona State University’s Biodesign Swette Center for Environmental Biotechnology, and researchers from the University of Arizona and Arizona State University, are participating in the study, “EFRI ELiS: Bioweathering Dynamics and Ecophysiology of Microbially Catalyzed Soil Genesis of Martian Regolith.” This National Science Foundation and NASA-funded project will use microorganisms from bacteria to remove perchlorates from Martian soil simulants and produce soil organic matter containing organic carbon and inorganic nutrients. Martian regolith contains high concentrations of toxic perchlorate salts that will impede plant cultivation in soil, jeopardizing food security and potentially causing health problems for humans, including cancer. Researchers will look at different bacterial populations and how well they are able to process and break down the perchlorates, as well as what kind of materials they produce when they do. They’ll also look at different temperatures and moisture conditions, as well as in the presence or absence of oxygen. Students in the Palmer Lab will receive the simulants after this process, try to replicate it, and then test how well the perchlorate-free regolith is able to grow plants. A challenge the researchers face is how they remove the toxic salts, as well as if they can remove all of them. Palmer cautioned that the possibility that removing the perchlorates does not necessarily mean the regolith is ready for farming. “You can’t make the cure worse than the disease, so we have to be ending up with regolith on the other side that’s better than when we started,” Palmer said. “We can’t trade perchlorates for some other toxic accumulating compound. Just because we’re removing the perchlorates doesn’t necessarily mean that we’re going to make the regolith better for plants. We might just make it not toxic anymore. How much does it improve is really what we’re trying to figure out.” Even without perchlorates, there are significant challenges to growing crops in Martian soil. While researchers have grown plants in simulated regolith, the regolith is not good for plant growth, as in addition to a lot of salts, it has a high pH and is very fine, which means it can ‘cement’ when wet, suffocating plant roots. Being able to grow in the soil instead of using hydroponics could also provide a more efficient, cost-effective solution. “There is always the option of hydroponic growth of food crops, but with a significant distance to Mars and the lack of readily available water, we need a different kind of plan,” said ASU’s Delgado. “If there is a possibility to grow plants directly in the soil, there are benefits in terms of water utilization and resources to get supplies to Mars.” Some of the microbial solutions the team is proposing could also help with studies of soils on Earth. “The best soils for agriculture on earth, they were taken up decades ago, and so now we’re trying to farm on new land that’s not really meant for agriculture, if you think about it,” Palmer said. “So, as we think about ways to convert it into better soil, I think this research helps teach us how to do that, but it also inspires.” The research will also allow Florida Tech students to get hands-on space agriculture experience. “We’re going to be training the grad students and the undergraduates who are going to be the researchers who take on those new challenges, so I think one of our most important products are going to be the students we train,” Palmer said. “We’ll deliver Mars soil, but we also deliver, I think, a future group of researchers.” If you're a reporter looking to know more about this topic - then let us help with your coverage. Dr. Andrew Palmer is an associate professor of biological sciences at Florida Tech and a go-to expert in the field of Martian farming. Andrew is available to speak with media regarding this and related topics. Simply click on his icon now to arrange an interview today.

Gene Editing Institute Travels to Salem for ‘Innovation Days’ Workshop
Education sessions bring CRISPR gene editing to high school students from diverse backgrounds Scientist-educators from ChristianaCare’s Gene Editing Institute held a workshop using CRISPR in a BoxTM at Salem Academy during Innovation Days in October at the school, located in Winston-Salem, North Carolina. These sessions followed a previous gene editing education workshop with Salem Academy students in January 2022. CRISPR in a BoxTM is a revolutionary toolkit that allows students to carry out a hands-on gene editing experiment while learning and analyzing the steps involved in a typical gene editing reaction. Scientists from the Gene Editing Institute also taught a condensed lesson about CRISPR gene editing’s utility in medicine and fielded questions from students about jobs in biotechnology, bioethics and sustainability in the lab. “It’s a really special opportunity that I know I wouldn’t get anywhere else,” said Mathilda Willenborg, a sophomore boarding student from Germany. “And I do feel like I’m learning a lot about gene editing that I definitely didn’t know before. The team makes it really easy and walks us through all the steps.” Last winter, Salem Academy became the first school in North Carolina to offer CRISPR in a Box as it pivoted its academic focus to STEAM (Science, Technology, Engineering, Art and Math). That first innovative workshop originated as a result of an idea from a ChristianaCare board member who attended Salem Academy. Gene Editing Institute Founder and Lead Scientist Eric Kmiec, Ph.D., made a virtual appearance as part of the latest sessions to encourage the students to pursue careers and pathways in biotechnology. “We’re so appreciative of our partnership with Salem Academy,” said Kmiec. “We want to take every chance we get to encourage more women to pursue careers in STEM. Women around the nation, and around the world, should have access to this groundbreaking technology, which will ultimately drastically change the way we treat and cure diseases. If we don’t have young women in that discussion, we’re missing out on valuable experiences and perspectives.” Salem Academy is the only all-female boarding and day high school on a college campus in the U.S. with a STEM focus. Women are achieving significant progress in STEM fields, representing 45% of students majoring in STEM, according to the Integrated Postsecondary Education Data System. However, women only represent 27% of STEM workers, with wide disparities in income in post-graduation employment. As of 2019, less than 30% of the world’s researchers were women, according to the UNESCO Institute for Statistics. The Gene Editing Institute commits to a mission of diversity and equity in its approach. This workshop reached 10 women, two of whom are international students. “Our ongoing partnership with the ChristianaCare Gene Editing Institute will help position our aspiring women scientists for future careers in biotechnology, science and medicine,” said Summer McGee, Ph.D., president of Salem Academy and College. “This is the type of experience that sets Salem Academy apart as a national leader in building the next generation of women leaders in STEAM.” The Gene Editing Institute itself is a national leader in female researchers. Women make up over 80% of scientists within the Institute and fill 75% of the principal investigator roles. The Institute pushes to address the gender gap and promote inclusivity through local outreach and state-spanning programs, like CRISPR in a Box. “We’re not here to do lip service,” said Brett Sansbury, Ph.D., principal investigator of the Discovery Branch of the Gene Editing Institute. “Too many companies make a plan or promise without any actionable steps. We’re taking those steps and bringing in opportunities for students who otherwise wouldn’t have had them.” To learn more about how to bring CRISPR in a Box to your school, visit https://geneeditinginstitute.com/products/education. About CRISPR in a BoxTM CRISPR in a BoxTM is the leading educational toolkit to teach gene editing. The exercise features a hands-on gene editing experiment, including a live readout within non-infectious E. coli bacteria. These experiments follow a gene editing reaction from beginning to end while teaching students the techniques scientists use to perform these reactions in real laboratory environments. CRISPR in a Box is distributed by Carolina Biological. To learn more, visit https://geneeditinginstitute.com/products/education.

Manuka honey could help to clear deadly drug-resistant lung infection – research
• Scientists develop a potential nebulisation treatment using manuka honey to clear a drug resistant lung infection that can be fatal in cystic fibrosis patients • Aston University researchers combined the antibiotic amikacin with manuka honey as a novel treatment for Mycobacterium abscessus • Using the manuka honey combination resulted in an eight-fold reduction in the dosage of the antibiotic A potential new treatment combining natural manuka honey with a widely used drug has been developed by scientists at Aston University to treat a potentially lethal lung infection and greatly reduce side effects of one of the current drugs used for its treatment. The findings, which are published in the journal Microbiology, show that the scientists in the Mycobacterial Research Group in the College of Health and Life Sciences at Aston University were able to combine manuka honey and the drug amikacin in a lab-based nebulisation formulation to treat the harmful bacterial lung infection Mycobacterium abscessus. Manuka honey is long known to have wide ranging medicinal properties, but more recently has been identified for its broad spectrum antimicrobial activity. Now scientists have found that manuka honey has the potential to kill a number of drug resistant bacterial infections such as Mycobacterium abscessus – which usually affects patients with cystic fibrosis (CF) or bronchiectasis. According to the Cystic Fibrosis Trust, CF is a genetic condition affecting around 10,800 people - one in every 2,500 babies born in the UK -and there are more than 100,000 people with the condition worldwide. The NHS defines bronchiectasis as a long-term condition where the airways of the lungs become widened, leading to a build-up of excess mucus that can make the lungs more vulnerable to infection.. In the study, the researchers used samples of the bacteria Mycobacterium abscessus taken from 16 infected CF patients. They then tested the antibiotic amikacin, combined with manuka honey, to discover what dosage was required to kill the bacteria. As part of the study the team used a lab-based lung model and nebuliser - a device that produces a fine spray of liquid often used for inhaling a medicinal drug. By nebulising manuka honey and amikacin together, it was found they could improve bacterial clearance, even when using lower doses of amikacin, which would result in less life-changing side-effects to the patient. In the UK, of the 10,800 people living with CF, Mycobacterium abscessus infects 13% of all patients with the condition. This new approach is advantageous not only because it has the potential to kill off a highly drug resistant infection, but because of the reduced side effects, benefitting quality of life and greatly improving survival chances for infected CF patients. Mycobacterium abscessus is a bacterial pathogen from the same family that causes tuberculosis, but this bug differs by causing serious lung infections in people (particularly children) with pre-existing lung conditions, such as CF and bronchiectasis, as well as causing skin and soft tissue infections. The bacteria is also highly drug resistant. Currently, patients are given a cocktail of antibiotics, consisting of 12 months or more of antimicrobial chemotherapy and often doesn’t result in a cure. The dosage of amikacin usually used on a patient to kill the infection is 16 micrograms per millilitre. But the researchers found that the new combination using manuka honey, required a dosage of just 2 micrograms per millitre of amikacin - resulting in a one eighth reduction in the dosage of the drug. Until now Mycobacterium abscessus has been virtually impossible to eradicate in people with cystic fibrosis. It can also be deadly if the patient requires a lung transplant because they are not eligible for surgery if the infection is present. Commenting on their findings, lead author and PhD researcher Victoria Nolan said: "So far treatment of Mycobacterium abscessus pulmonary infections can be problematic due to its drug resistant nature. The variety of antibiotics required to combat infection result in severe side effects. "However, the use of this potential treatment combining amikacin and manuka honey shows great promise as an improved therapy for these terrible pulmonary infections. “There is a need for better treatment outcomes and in the future we hope that this potential treatment can be tested further.” Dr Jonathan Cox, senior lecturer in microbiology, Aston University said: “By combining a totally natural ingredient such as manuka honey with amikacin, one of the most important yet toxic drugs used for treating Mycobacterium abscessus, we have found a way to potentially kill off these bacteria with eight times less drug than before. This has the potential to significantly reduce amikacin-associated hearing loss and greatly improve the quality of life of so many patients – particularly those with cystic fibrosis. “I am delighted with the outcome of this research because it paves the way for future experiments and we hope that with funding we can move towards clinical trials that could result in a change in strategy for the treatment of this debilitating infection.” Dr Peter Cotgreave, chief executive of the Microbiology Society said: "The Microbiology Society is proud to support the scientific community as it explores innovative solutions to overcome the growing global challenge of antimicrobial resistance. This study demonstrates one of many ways in which microbiologists are pioneering new methods to tackle drug-resistant infections, by incorporating natural products, like manuka honey, into existing therapies." For more information about the School of Biosciences, please visit our website.

Meet the astrobiologist and her students who are searching for life on Mars
By Emma Richards, University of Florida From a young age, Amy Williams wondered if life existed beyond Earth amidst the dark abyss of space, stars and planets — a curiosity that years later landed her a career researching and exploring Mars. Williams, an assistant professor of geology and an astrobiologist at the University of Florida, works as a participating scientist on the Perseverance and Curiosity Rover Science Teams and previously served as a postdoctoral research associate at NASA’s Goddard Space Flight Center. As an astrobiologist and geobiologist, she uses techniques from geology, microbiology and chemistry to search for life beyond Earth. “Even as a little kid watching meteor showers with my family, I wondered if there was someone out there in the stars looking back at Earth.” “Even as a little kid watching meteor showers with my family, I wondered if there was someone out there in the stars looking back at Earth,” she said in an episode of the From Florida podcast. “It’s been a passion of mine my whole career and now it’s the most amazing opportunity to serve on both of the active Mars rover missions.” Williams’ journey to Mars began as a graduate student when a research professor gave her the opportunity to work on the NASA Curiosity mission. From there, Williams built her way up and is now a participating scientist working on day-to-day rover operations. Williams also is opening doors for graduate students at UF to work on Mars research, helping upcoming generation of scientists follow her path. She is specifically interested in involving women and underrepresented groups in her work. Based on her research, Williams said life on Mars, if found, will likely look less like Marvin the Martian and more like microbial life similar to bacteria on Earth. Curiosity landed on Mars in 2012 and Perseverance landed in 2021. The rovers are searching for potential life on Mars by going to habitable environments and searching for evidence of water and essential elements that could supported such life forms. Curiosity has spent nearly its entire mission exploring a large five-kilometer-tall mountain in Gale Crater known as Mount Sharp. The scientists can see Mars’ history and climate based on changes in the chemistry and sediments of the mountain. As for Perseverance, the rover is exploring Jezero Crater, with emphasis on its delta, a geologic deposit that is formed when water from a river flows into a lake. Perseverance will help collect rock and sediment samples from Mars that will be the first brought back to Earth. NASA is also working on a program to eventually send humans to Mars, which will likely take many decades; the first stage in the project will be returning humans to the Moon. “But in the meanwhile, these robots, these rovers that we send to the red planet, they are our proxy,” Williams said. “And looking through the robot rover’s eyes, the images that are returned to us, I recognize this is the closest I will ever be to standing on Mars and looking up at these beautiful geological units, looking up at an alien world that’s so familiar because the tenets of geology apply on Mars, the same as they do on Earth.” To hear more about the Amy Williams' Mars research, listen to the episode on From Florida at this link. Listen to other episodes in the "From Florida" series at this link. To learn more about her work, watch this video featuring Professor Williams:

Sweeteners may be linked to increased cancer risk – new research
Sweeteners have long been suggested to be bad for our health. Studies have linked consuming too many sweeteners with conditions such as obesity, type 2 diabetes and cardiovascular disease. But links with cancer have been less certain. An artificial sweetener, called cyclamate, that was sold in the US in the 1970s was shown to increase bladder cancer in rats. However, human physiology is very different from rats, and observational studies failed to find a link between the sweetener and cancer risk in humans. Despite this, the media continued to report a link between sweeteners and cancer. But now, a study published in PLOS Medicine which looked at over 100,000 people, has shown that those who consume high levels of some sweeteners have a small increase in their risk of developing certain types of cancer. To assess their intake of artificial sweeteners, the researchers asked the participants to keep a food diary. Around half of the participants were followed for more than eight years. The study reported that aspartame and acesulfame K, in particular, were associated with increased cancer risk – especially breast and obesity-related cancers, such as colorectal, stomach and prostate cancers. This suggests that removing some types of sweeteners from your diet may reduce the risk of cancer. Cancer risk Many common foods contain sweeteners. These food additives mimic the effect of sugar on our taste receptors, providing intense sweetness with no or very few calories. Some sweeteners occur naturally (such as stevia or yacon syrup). Others, such as aspartame, are artificial. Although they have few or no calories, sweeteners still have an effect on our health. For example, aspartame turns into formaldehyde (a known carcinogen) when the body digests it. This could potentially see it accumulate in cells and cause them to become cancerous. Our cells are hard-wired to self-destruct when they become cancerous. But aspartame has been shown to “switch off” the genes that tell cancer cells to do this. Other sweeteners, including sucralose and saccharin, have also been shown to damage DNA, which can lead to cancer. But this has only been shown in cells in a dish rather than in a living organism. Sweeteners can also have a profound effect on the bacteria that live in our gut. Changing the bacteria in the gut can impair the immune system, which could mean they no longer identify and remove cancerous cells. But it’s still unclear from these animal and cell-based experiments precisely how sweeteners initiate or support cancerous changes to cells. Many of these experiments would also be difficult to apply to humans because the amount of sweetener was given at much higher doses than a human would ever consume. The results from previous research studies are limited, largely because most studies on this subject have only observed the effect of consuming sweeteners without comparing against a group that hasn’t consumed any sweeteners. A recent systematic review of almost 600,000 participants even concluded there was limited evidence to suggest heavy consumption of artificial sweeteners may increase the risk of certain cancers. A review in the BMJ came to a similar conclusion. Although the findings of this recent study certainly warrant further research, it’s important to acknowledge the study’s limitations. First, food diaries can be unreliable because people aren’t always honest about what they eat or they may forget what they have consumed. Although this study collected food diaries every six months, there’s still a risk people weren’t always accurately recording what they were eating and drinking. Though the researchers partially mitigated this risk by having participants take photos of the food they ate, people still might not have included all the foods they ate. Based on current evidence, it’s generally agreed that using artificial sweeteners is associated with increased body weight – though researchers aren’t quite certain whether sweeteners directly cause this to happen. Although this recent study took people’s body mass index into account, it’s possible that changes in body fat may have contributed to the development of many of these types of cancers – not necessarily the sweeteners themselves. Finally, the risk of developing cancer in those who consumed the highest levels of artificial sweeteners compared with those who consumed the lowest amounts was modest – with only at 13% higher relative risk of developing cancer in the study period. So although people who consumed the highest amounts of sweetener had an increased risk of developing cancer, this was still only slightly higher than those with the lowest intake. While the link between sweetener use and diseases, including cancer, is still controversial, it’s important to note that not all sweeteners are equal. While sweeteners such as aspartame and saccharin may be associated with ill health, not all sweeteners are. Stevia, produced from the Stevia rebaudiana plant, has been reported to be useful in controlling diabetes and body weight, and may also lower blood pressure. The naturally occurring sugar alcohol, xylitol, may also support the immune system and digestion. Both stevia and xylitol have also been shown to protect from tooth decay, possibly because they kill bad oral bacteria. So the important choice may be not the amount of sweetener you eat but the type you use.

In new study in journal Gene Therapy, researchers at ChristianaCare’s Gene Editing Institute describe how the advance is validating the safety and efficacy of their novel approach for using CRISPR to improve lung cancer treatments A new study from scientists at ChristianaCare’s Gene Editing Institute is advancing the safety and efficacy of using CRISPR gene editing in patient treatments by demonstrating how to identify and evaluate the broad-based biological impact of gene editing on targeted tissues, where the edits are designed to fully disable or “knock out” a specific sequence of genetic code. The work, published today in the Nature journal Gene Therapy, supports the Institute’s efforts to improve lung cancer treatments by using CRISPR to disable or alter a master regulator gene to prevent it from producing a protein that blunts the impact of chemotherapy. “We found that when you use CRISPR, the edits sometimes end up altering rather than completely disabling the target gene, so we developed a process to gain a more complete understanding of what that means for patients,” said Eric Kmiec, Ph.D., executive director and chief scientific officer of ChristianaCare’s Gene Editing Institute and the principal author of the study. Dr. Kmiec said that for his team’s lung cancer work, “We discovered that even when our CRISPR-based genetic manipulation did not completely disable the targeted gene, it altered it in ways that appear to make lung cancer tumors more sensitive to chemotherapy. Validating lung cancer research using CRISPR “We were fortunate that our strategy for using CRISPR to improve lung cancer treatments has been validated once again,” he added. “But our commitment to conducting an unbiased assessment of our approach highlights the importance of examining all potential outcomes of an attempt to use CRISPR to knock out a specific gene. Specifically, anyone developing CRISPR therapies needs to be on the lookout for edits that don’t fully knock out a section of DNA code—and evaluate the potential impacts for patients. They could be positive, as they were in our case, negative or neutral, but they need to be known.” Much of the excitement around medical applications of CRISPR involves using the tool to disable harmful genes by editing or “knocking out” a specific sequence of DNA code. But there is increasing evidence that in the wake of a CRISPR edit, cells may remain that contain merely an altered form of the targeted code that allows the gene to continue to produce biologically active proteins. Scientists at the Gene Editing Institute are investigating the potential of using CRISPR to disable a gene called NRF2 to alter production of the protein that protects squamous cell carcinoma lung cancer tumors from the effects of chemotherapy or radiation. They already have shown, in studies with tumor cells and in animals, that they can selectively target the NRF2 gene without affecting normal cells, where the gene confers health benefits. In the present study they wanted to go further. They wanted to fully understand the implications of a CRISPR gene edit that allowed the NRF2 gene to retain enough DNA code to continue making a version of the protein, albeit in an altered or truncated form. The team is laying the groundwork for a clinical trial that would use CRISPR to improve the efficacy of conventional chemotherapy and radiation treatments. Dr. Kmiec said that before proceeding, he wanted his team to develop a clear process for identifying and evaluating all outcomes of CRISPR edits. Identifying and understanding the diversity of genetic outcomes produced by CRISPR-directed gene editing has been a centerpiece of the foundational research programs established by the Gene Editing Institute. Using CRISPR in a safe way “We carry out experiments in an unbiased fashion, not hoping for a particular outcome, but with patient safety and efficacy serving as the true north for our scientific endeavors,” Dr. Kmiec said. “No matter what we uncover or elucidate, the insights will help both ChristianaCare and the entire field use CRISPR in a safer and more efficacious manner.” The researchers found multiple cells where the targeted strand of DNA code in the NRF2 gene was not completely knocked out. Rather, following the CRISPR edit, cells emerged that had retained enough of the original code to continue producing a different form of the protein. Tests revealed that cancer tumor cells generating these altered proteins may be more vulnerable to chemotherapy drugs. "For the work we are doing with NRF2, the truncated proteins generated by the CRISPR edit appear to be beneficial for making tumors more sensitive to treatment,” said lead author Kelly Banas, Ph.D. “But the key point is these proteins were clearly biologically active. And that means we needed to determine their potential impact on the safety and efficacy of using CRISPR to treat lung cancer patients.” Dr. Banas noted that the study points to the limits of considering a CRISPR edit to be successful simply by testing for the absence of a targeted protein in its original form. She said by that standard, their edit was successful. The edited NRF2 genes were no longer producing the same protein. But she said if that’s all the ChristianaCare team had looked for, they would have missed the altered proteins coming from the NRF2 gene—and overlooked an important outcome that, in this case, strengthens the original hypothesis and experimental approach: that using CRISPR to target the NRF2 gene holds promise for improving outcomes for lung cancer patients. Importance of due diligence “The process we describe in this study is a template that should be followed in any effort to develop CRISPR as a medical treatment,” Dr. Kmiec said. “We’re part of a health care organization where patient safety is the top priority. We also are working at the vanguard of an exciting area of cutting-edge medicine, where a failure to conduct due diligence could cause tragic outcomes that would set back this field for decades. With this study, we have validated a process that can help this field move forward rapidly but safely.” CRISPR stands for “clustered regularly interspaced short palindromic repeats.” It is a defense mechanism found in bacteria that can recognize and slice up the DNA of invading viruses. Scientists have learned how to modify this mechanism so it can be directed to “edit” specific sequences of DNA code. About ChristianaCare’s Gene Editing Institute The Gene Editing Institute, a worldwide leader in CRISPR gene editing technology and the only institute of its kind based within a community health care system, takes a patient-first approach in all its research to improve the lives of people with life-threatening disease. Since 2015, researchers at the Gene Editing Institute have been involved in several ground-breaking firsts in the field, including the development of the first CRISPR gene editing tool to allow DNA repairs outside the human cell which will rapidly speed therapies to patients and the ExACT ™pathway of single-stranded DNA repair, which increased the on-target efficacy of CRISPR and paved the way for new CRISPR breakthroughs in precise DNA edits. Its researchers created CRISPR in a Box™, the leading educational toolkit to teach gene editing, DECODR™, recognized as the most user-friendly and precise analytical tool to understand the diversity of genetic outcomes of gene editing and are currently developing a patient trial for lung cancer using CRISPR.

Antimicrobial resistance now causes more deaths than HIV/AIDS and malaria worldwide – new study
Antimicrobial resistance is spreading rapidly worldwide, and has even been likened to the next pandemic – one that many people may not even be aware is happening. A recent paper, published in Lancet, has revealed that antimicrobial resistant infections caused 1.27 millions deaths and were associated with 4.95 million deaths in 2019. This is greater than the number of people who died from HIV/AIDS and malaria that year combined. Antimicobial resistance happens when infection-causing microbes (such as bacteria, viruses or fungi) evolve to become resistant to the drug designed to kill them. This means than an antibiotic will no longer work to treat that infection anymore. The new findings makes it clear that antimicrobial resistance is progressing faster than the previous worst-case scenario estimates – which is of concern for everyone. The simple fact is that we’re running out of antibiotics that work. This could mean everyday bacterial infections become life-threatening again. While antimicrobial resistance has been a problem since penicillin was discovered in 1928, our continued exposure to antibiotics has enabled bacteria and other pathogens to evolve powerful resistance. In some cases, these microbes are resistant even to multiple different drugs. This latest study now shows the current scale of this problem globally – and the harm it’s causing. Global problem The study involved 204 countries around the world, looking at data from 471 million individual patient records. By looking at deaths due to and associated with antimicrobial resistance, the team was then able to estimate the impact antimicrobial resistance had in each country. Antimicrobial resistance was directly responsible for an estimated 1.27 million deaths worldwide and was associated with an estimated 4.95 millions deaths. In comparison, HIV/AIDS and malaria were estimated to have caused 860,000 and 640,000 deaths respectively the same year. The researchers also found that low- and middle-income countries were worst hit by antimicrobial resistance – although higher income countries also face alarmingly high levels. They also found that of the 23 different types of bacteria studied, drug resistance in only six types of bacteria contributed to 3.57 million deaths. The report also shows that 70% of deaths that resulted from antimicrobial resistance were caused by resistance to antibiotics often considered the first line of defence against severe infections. These included beta-lactams and fluoroquinolones, which are commonly prescribed for many infections, such as urinary tract, upper- and lower-respiratory and bone and joint infections. This study highlights a very clear message that global antimicrobial resistance could make everyday bacterial infections untreatable. By some estimates, antimicrobial resistance could cause 10 million deaths per year by 2050. This would overtake cancer as a leading cause of death worldwide. Next pandemic Bacteria can develop antimicrobial resistance in a number of ways. First, bacteria develop antimicrobial resistance naturally. It’s part of the normal push and pull observed throughout the natural world. As we get stronger, bacteria will get stronger too. It’s part of our co-evolution with bacteria – they’re just quicker at evolving than we are, partly because they replicate faster and get more genetic mutations than we do. But the way we use antibiotics can also cause resistance. For example, one common cause is if people fail to complete a course of antibiotics. Although people may feel better a few days after starting antibiotics, not all bacteria are made equal. Some may be slower to be affected by the antibiotic than others. This means that if you stop taking the antibiotic early, the bacteria that were initially able to avoid the effect of the antibiotics will be able to multiply, thus passing their resistance on. Likewise, taking antibiotics unnecessarily can help bacteria to evolve resistance to antibiotics faster. This is why it’s important not to take antibiotics unless they’re prescribed, and to only use them for the infection they’re prescribed for. Resistance can also be spread from person to person. For example, if someone who has antibiotic-resistant bacteria in their nose sneezes or coughs, it may be spread to people nearby. Research also shows that antimicrobial resistance can be spread through the environment, such as in unclean drinking water. The causes driving this global antimicrobial resistance crisis are complex. Everything from how we take antibiotics to environmental pollution with antimicrobial chemicals, use of antibiotics in agriculture and even preservatives in our shampoo and toothpaste are all contributing to resistance. This is why a global, unified effort will be needed to make a difference. Urgent change is needed in many industries to slow the spread of antimicrobial resistance. Of the greatest importance is using the antibiotics we have smarter. Combination therapy could hold the answer to slowing down antimicrobial resistance. This involves using several drugs in combination, rather than one drug on its own – making it more difficult for bacteria to evolve resistance, while still successfully treating an infection. The next pandemic is already here – so further investment in research that looks at how we can stop this problem will be key.




